This web page was produced as an assignment for genetics 564, an undergraduate course at UW-Madison
Model Organisms
Model organisms have substantial utility in the research setting (5). A model organism is a plant, animal or microbe that can be used to study a biological process. Since all organisms have derived from a common ancestor, certain organisms have retained certain processes and principles that can be studied and provide insight into other organisms such as humans. This has incredible power due to the fact studying disease processes and performing experiments on people are typically unethical. Thus, studying other organisms that possess similar biological processes can provide insight into understanding the cause of a disease. This also aids in seeking out potential diagnostic tests and treatments. Model organisms have been incredibly useful for understanding complex phenomenon in humans.
Model organisms have substantial utility in the research setting (5). A model organism is a plant, animal or microbe that can be used to study a biological process. Since all organisms have derived from a common ancestor, certain organisms have retained certain processes and principles that can be studied and provide insight into other organisms such as humans. This has incredible power due to the fact studying disease processes and performing experiments on people are typically unethical. Thus, studying other organisms that possess similar biological processes can provide insight into understanding the cause of a disease. This also aids in seeking out potential diagnostic tests and treatments. Model organisms have been incredibly useful for understanding complex phenomenon in humans.
RNA Interference
RNAi techniques utilize naturally occurring processes of some organisms for research purposes (8,9). RNA interference, otherwise known as RNAi, can silence specific gene transcripts, meaning it can destroy a transcript prior to translation (8,9). This is a powerful tool in the research field because you can study and mimic disease states that involve the loss of function of a certain protein product and prevent that protein expression in a model organism. RNAi is also a promising method in research for therapies for diseases. Diseases that would benefit from therapeutic RNAi methods are those that have over expression of certain transcripts or abnormal pathological protein products. For example, if a disease is caused by over expression of certain genes, it is believed these bad transcripts can be targeted and destroyed prior to being translated into proteins. RNAi techniques can be engineered to target specific transcripts of interest. Therefore, this technique can be used to shut off the expression of mRNAs, and consequently, specific proteins. This can be taken advantage of in a single celled embryo to make a knockout organism. A video visually depicting how RNAi works is shown below.
RNAi techniques utilize naturally occurring processes of some organisms for research purposes (8,9). RNA interference, otherwise known as RNAi, can silence specific gene transcripts, meaning it can destroy a transcript prior to translation (8,9). This is a powerful tool in the research field because you can study and mimic disease states that involve the loss of function of a certain protein product and prevent that protein expression in a model organism. RNAi is also a promising method in research for therapies for diseases. Diseases that would benefit from therapeutic RNAi methods are those that have over expression of certain transcripts or abnormal pathological protein products. For example, if a disease is caused by over expression of certain genes, it is believed these bad transcripts can be targeted and destroyed prior to being translated into proteins. RNAi techniques can be engineered to target specific transcripts of interest. Therefore, this technique can be used to shut off the expression of mRNAs, and consequently, specific proteins. This can be taken advantage of in a single celled embryo to make a knockout organism. A video visually depicting how RNAi works is shown below.
ACVR1 Mutant Phenotypes in Model Organisms
Caenorhabditis elegans (nematode)
|
From the protein homology search, daf-1 was found to be 40% similar to ACVR1 in humans. Using wormbase, daf-1 does indeed code for a TGF-beta receptor similarly to ACVR1 in humans, however, the website states that there is no human disease data for daf-1. The absence of data for FOP may be because there is not enough research on it or that the gene is too dissimilar in the nematode and does not involve similar biological outcomes as seen in FOP. Mutations in the daf-1 serine threonine kinase protein results in constitutive formation of dauer larvae even under optimal conditions.
|
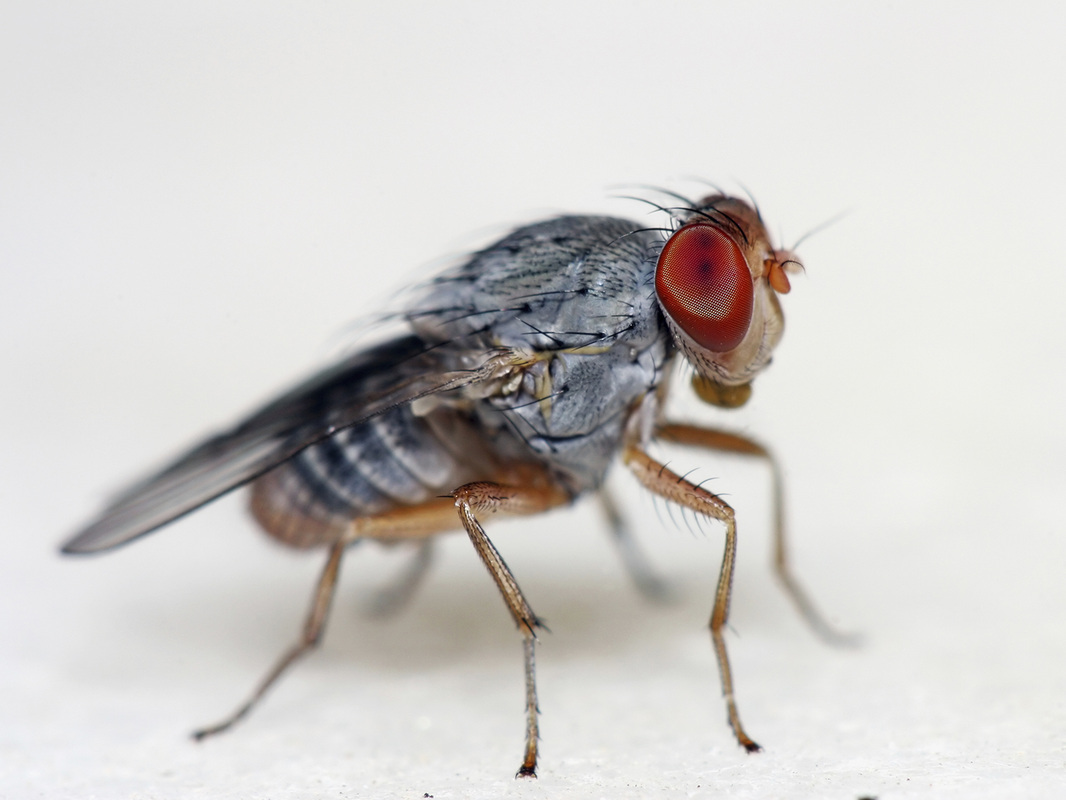
Drosophila melanogaster (fruit fly)
|
Using a database called Flybase, information about homologs in flies can be investigated. Fruit flies have a protein homolog to human ACVR1 called sax. This protein is also a TGF-beta protein kinase involved in signaling. This protein is 51% identical to the human homolog. Using the gene ontology found on this database, it appears that this protein is involved in different biological processes and outcomes than in humans. Additionally, it appears that so far there is no information on human disease model data yet for this protein.
|
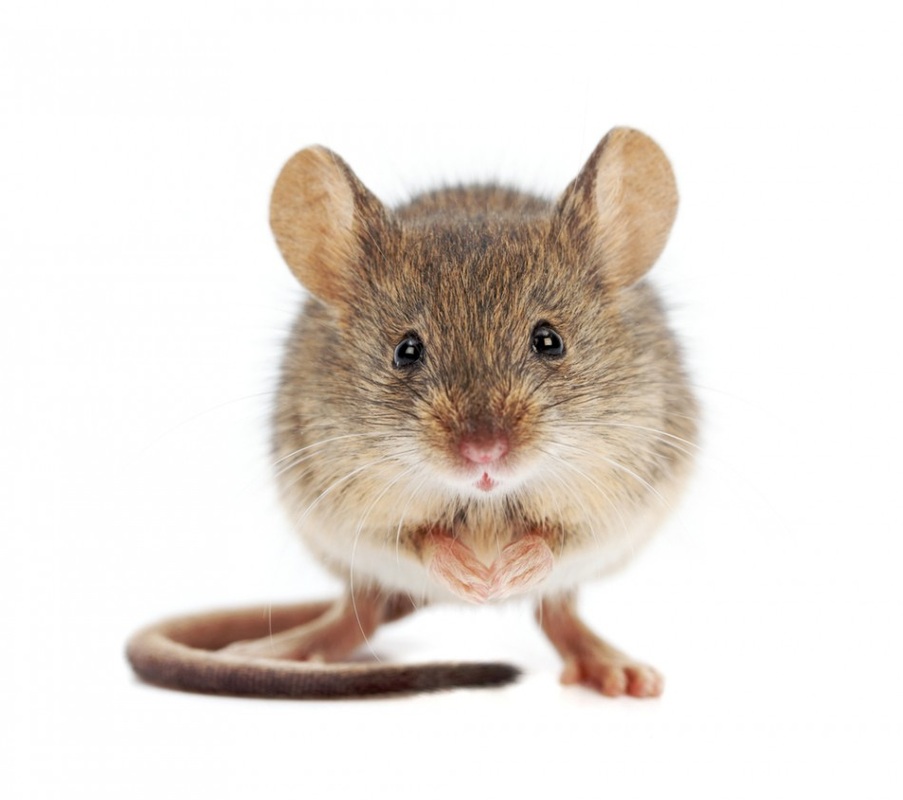
Mus musculus (mouse)
|
Using MGI, which is a mouse database, mouse models have been made to study FOP. One model was made by creating a transgenic mouse while the other was created by making a targeted mutation. Because FOP is not characterized by a loss of function of the protein, producing a knockout mouse would not adequetly simulate the disase phenotype in humans. In human FOP, the signaling is excessive as opposed to being nonfunctional, thus, it makes sense why these transgenic mice for this specific amino acid change were created. It appears the mutant mice strains possess the disease phenotypes that are seen in humans. Thus, using a mouse model appears to be a useful model for FOP research.
|
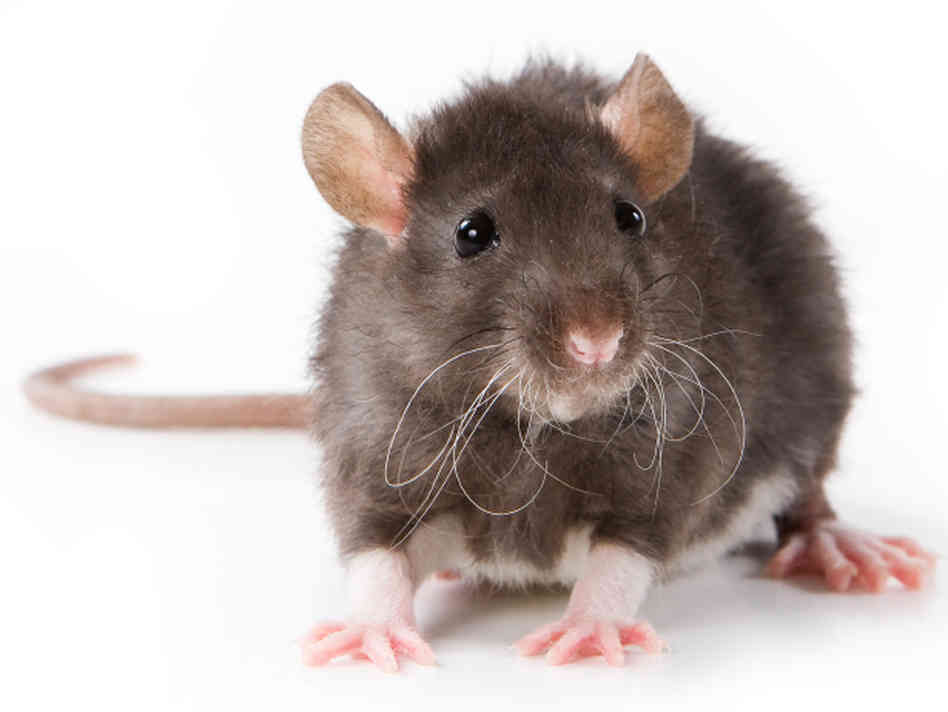
Rattus rattus (rat)
|
The RGD database has associated with ACVR1 to myositis ossificans, otherwise referred to FOP. Therefore, this indicates rats are also a suitable model organism. Various strain sequence variants have been created for rats.
|
Analysis:
Due to the fact no protein homologs were found in microbes or plants, the data for these websites were not included but can be found at Dictybase, TAIR, SDG, and Pombase. From these databases, it can be seen that rodents such as mice and rats may be the optimal model organism to pursue for FOP research. First, these protein homologes in rats and mice are very similar (97 percent and 98 percent identical) to the ACVR1 in humans and perform the same biological processes and signaling pathways. Secondly, these organisms have a defined muscular and skeletal system, making it easy to visualize the disease phenotypes in these model organisms. Moreover, the wormbase and flybase indicate with gene ontology that this gene is involved in other biological processes than ACVR1 in humans, therefore making it not the optimal model. RNAi or CRISPR/Cas9 would not be a useful technique in creating a knockout model organism due to the fact that in FOP, the protein is still functional; it just has excessive activity. Therefore, a knockout model would not adequately reflect the molecular mechanisms causing the phenotype. This explains why transgenic and targeted mutation strains were created to study FOP in rodents. Interestingly, because RNAi does destroy transcripts for a certain mRNA, this technique has been sought out as a potential treatment for FOP. You would expect excessive gene expression of downstream targets in the BMP pathway, thus, if RNAi can limit the expression of these unnecessary transcripts, it can be hypothesized that this may decrease the severity of the disease by controlling flare-ups of bone growth. This is an area of further research (10). FOP is an autosomal dominant disease, so research is being performed to see if RNAi can knockout the mutant copy of ACVR1, allowing the normal copy of ACVR1 to create the appropriate protein. Therefore, gene therapy is an area of research interest.
Due to the fact no protein homologs were found in microbes or plants, the data for these websites were not included but can be found at Dictybase, TAIR, SDG, and Pombase. From these databases, it can be seen that rodents such as mice and rats may be the optimal model organism to pursue for FOP research. First, these protein homologes in rats and mice are very similar (97 percent and 98 percent identical) to the ACVR1 in humans and perform the same biological processes and signaling pathways. Secondly, these organisms have a defined muscular and skeletal system, making it easy to visualize the disease phenotypes in these model organisms. Moreover, the wormbase and flybase indicate with gene ontology that this gene is involved in other biological processes than ACVR1 in humans, therefore making it not the optimal model. RNAi or CRISPR/Cas9 would not be a useful technique in creating a knockout model organism due to the fact that in FOP, the protein is still functional; it just has excessive activity. Therefore, a knockout model would not adequately reflect the molecular mechanisms causing the phenotype. This explains why transgenic and targeted mutation strains were created to study FOP in rodents. Interestingly, because RNAi does destroy transcripts for a certain mRNA, this technique has been sought out as a potential treatment for FOP. You would expect excessive gene expression of downstream targets in the BMP pathway, thus, if RNAi can limit the expression of these unnecessary transcripts, it can be hypothesized that this may decrease the severity of the disease by controlling flare-ups of bone growth. This is an area of further research (10). FOP is an autosomal dominant disease, so research is being performed to see if RNAi can knockout the mutant copy of ACVR1, allowing the normal copy of ACVR1 to create the appropriate protein. Therefore, gene therapy is an area of research interest.
Gene Analysis and Transcriptome Analysis
What are Transcripts?
How is it all the cells in an individual contain the same DNA but are vastly different? In a human body, our cells can take the identity of multiple cell types such as neurons, white blood cells, muscle cells, bone cells, skin cells, etc. The key to understanding this phenomenon is in studying gene expression. Differences in gene expression result in different protein products with different activities in a cell. These differences in gene expression account for the vast variety of cell types and consequently, cellular functions. However, disease states can alter the gene expression pattern in a certain cell type. Different diseases may alter the expression of a gene or multiple genes. This is where transcriptomics comes in. Transcripts are the messenger RNA in cells. By studying mRNA, this gives insight into which genes are being actively translated. By comparing the expression of the pathological and the normal state, insight into the differential gene expression can be found. This differential gene expression can provide clues as to the molecular mechanisms involved in the transdifferentiation of soft tissue into bone in FOP. Below outlines ways in which transcripts and cell expression can be studied.
What are GEO profiles?
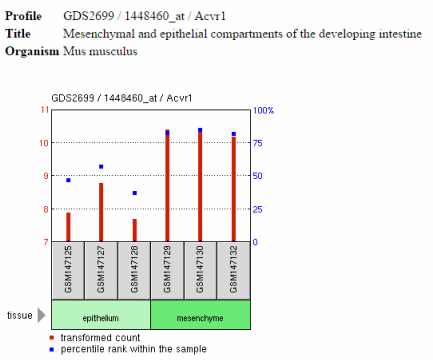
Gene Expression Omnibus or GEO Profiles is a database that contains information on gene expression profiles in the form of a chart (10). The charts illustrate the expression of a specific gene across all the samples in a data set. These charts are derived from various experiments. This is useful because it allows for a researcher to see if the gene is deferentially expressed under various conditions. The image below represents an example of a GEO profile for ACVR1. This image illustrates the expression of ACVR1 in epithelium compartments compared to the mesenchymal cells in the intestines during mouse development. This is one example from the 4,868 results that were found to be associated with ACVR1.
What are Transcripts?
How is it all the cells in an individual contain the same DNA but are vastly different? In a human body, our cells can take the identity of multiple cell types such as neurons, white blood cells, muscle cells, bone cells, skin cells, etc. The key to understanding this phenomenon is in studying gene expression. Differences in gene expression result in different protein products with different activities in a cell. These differences in gene expression account for the vast variety of cell types and consequently, cellular functions. However, disease states can alter the gene expression pattern in a certain cell type. Different diseases may alter the expression of a gene or multiple genes. This is where transcriptomics comes in. Transcripts are the messenger RNA in cells. By studying mRNA, this gives insight into which genes are being actively translated. By comparing the expression of the pathological and the normal state, insight into the differential gene expression can be found. This differential gene expression can provide clues as to the molecular mechanisms involved in the transdifferentiation of soft tissue into bone in FOP. Below outlines ways in which transcripts and cell expression can be studied.
What are GEO profiles?
Gene Expression Omnibus or GEO Profiles is a database that contains information on gene expression profiles in the form of a chart (10). The charts illustrate the expression of a specific gene across all the samples in a data set. These charts are derived from various experiments. This is useful because it allows for a researcher to see if the gene is deferentially expressed under various conditions. The image below represents an example of a GEO profile for ACVR1. This image illustrates the expression of ACVR1 in epithelium compartments compared to the mesenchymal cells in the intestines during mouse development. This is one example from the 4,868 results that were found to be associated with ACVR1.
|
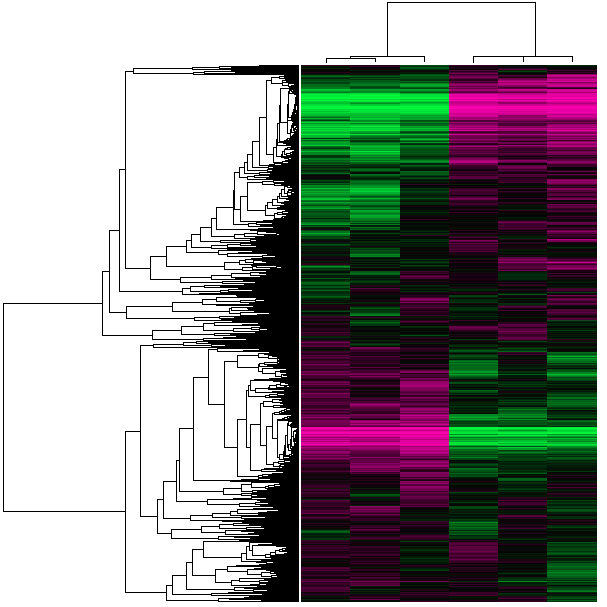
What are GEO DataSets? GEO datasets allow for a visual representation of the expression pattern found in two conditions. Based on the colors, it can be determined if the expression in the experiment is similar or different to the standard. No cluster heat maps were available to compare FOP muscle tissue gene expression to the wildtype state. This would be an interesting experiment to pursue. Since no heat maps have been performed specifically for FOP, therefore, no AMIGO/GO results were available. What pathway is involved?
Using a website called BioCarta, images of pathways involved with certain biological processes can be searched. The bone morphogenetic pathway is not currently listed on the website. Analysis:
Although these research tools are extremely informative, no cluster heat maps or GEO profiles have been explicitly performed regarding FOP. This would be a useful test to perform in order to see the differential expression in the disease state compared to the normal state. Although many GEO profiles were present for ACVR1, this protein has numerous activities and is involved in a substantial amount of biological activities. Understanding the pathways and differential expression in the disease state is an important area for future FOP research, especially in understanding how this metamorphosis between cell types occurs in FOP. |
Chemical Genetics
What is chemical genetics?
Chemical genetics involves screening for small molecules that bind to a particular protein (1,3). These small molecules have important utility in drug discovery because they can alter the activity of a protein. A well known example of the power of small molecules is the drug ibuprofen. Ibuprofen is a well known small molecule used to help relieve pain. This molecule diminishes headaches by going into our bloodstream and binding to an enzyme that is involved in the synthesis of the pain signaling molecules. The binding of the small molecule affects the capacity of the enzyme to perform its normal function. A video illustrating this can be found below.
What is chemical genetics?
Chemical genetics involves screening for small molecules that bind to a particular protein (1,3). These small molecules have important utility in drug discovery because they can alter the activity of a protein. A well known example of the power of small molecules is the drug ibuprofen. Ibuprofen is a well known small molecule used to help relieve pain. This molecule diminishes headaches by going into our bloodstream and binding to an enzyme that is involved in the synthesis of the pain signaling molecules. The binding of the small molecule affects the capacity of the enzyme to perform its normal function. A video illustrating this can be found below.
What small molecules interact with ACVR1?
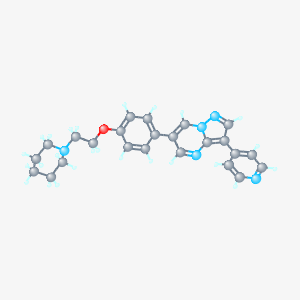
Small molecules play an integral role in pharmaceutical companies for finding drugs. It is the hope that small molecules can be a potential vehicle to treating FOP. In FOP, research has shown that the signaling pathway that forms bone is over stimulated due to a mutation in the TGF-beta motif. This mutation causes the inhibitor to be unable to bind to the TGF-beta binding domain when signaling is not present. Since this inhibitor is no longer able to regulate this pathway, the pathway is constitutively active, thus leading to the disease phenotype of excessive bone growth at soft tissue sites. Moreover, it is hypothesized that by finding a small molecule to inhibit the aberrant signaling, perhaps this will prevent or limit the amount of excessive bone growth. Chemical libraries contain all the small molecules that interact with a specific protein. There are a few databases that can be used such as Pubchem and Chembank. However, no results were found on Chembank. Using the target function on Pubchem, the list of chemicals that interact with ACVR1 is listed in the document below. These small molecules may be a direction of future drug discovery and research for FOP.
| pubchemACVR1smallmolecules | |
| File Size: | 67 kb |
| File Type: | csv |
Analysis:
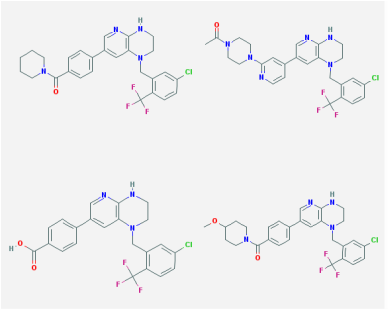
By the nature of FOP, small molecules are a promising avenue to explore for potential drug treatments. FOP is caused by a single missense mutation in the highly conserved TGF-beta motif, and consequently results in unnecessary BMP pathway signaling. Therefore, small molecules that act as inhibitors have been the prime focus of therapies for FOP. Using Pubchem, 144 small molecules were found. The activities of these small molecules varied ranging from binding constants for ACVR1 kinase domains, competitive binding affinity to ACVR1, inhibitors of ACVR1, increased binding affinity to human ALK-2, inhibitors of ALK-2, BMP inhibitors and kinase inhibitors. Images of some of the chemical structures of the inhibitors are shown above. Many of these molecules are large aromatic molecules. Image 6 illustrates a small molecule called Dorsmorphin (4). This small molecule is an inhibitor of ACVR1 and was the focus of drug therapy research for FOP. Using these databases to discover small molecules can be an invaluable tool for drug development and is especially applicable for FOP research. Therefore, further research on the effectiveness of these inhibitors should be tested in a FOP model organism. These small molecules studies can help elucidate ways to diminish the excessive BMP pathway signaling in the pathological state.
By the nature of FOP, small molecules are a promising avenue to explore for potential drug treatments. FOP is caused by a single missense mutation in the highly conserved TGF-beta motif, and consequently results in unnecessary BMP pathway signaling. Therefore, small molecules that act as inhibitors have been the prime focus of therapies for FOP. Using Pubchem, 144 small molecules were found. The activities of these small molecules varied ranging from binding constants for ACVR1 kinase domains, competitive binding affinity to ACVR1, inhibitors of ACVR1, increased binding affinity to human ALK-2, inhibitors of ALK-2, BMP inhibitors and kinase inhibitors. Images of some of the chemical structures of the inhibitors are shown above. Many of these molecules are large aromatic molecules. Image 6 illustrates a small molecule called Dorsmorphin (4). This small molecule is an inhibitor of ACVR1 and was the focus of drug therapy research for FOP. Using these databases to discover small molecules can be an invaluable tool for drug development and is especially applicable for FOP research. Therefore, further research on the effectiveness of these inhibitors should be tested in a FOP model organism. These small molecules studies can help elucidate ways to diminish the excessive BMP pathway signaling in the pathological state.
Image References:
1. http://cdn.isciencetimes.com/data/images/full/2014/05/16/6508-mouse.jpg
2. http://content62.eol.org/content/2012/09/26/23/54778_orig.jpg
3. http://www.redorbit.com/media/uploads/2012/02/science-021712-001-617x416.jpg
4. http://arlingtonva.s3.amazonaws.com/wp-content/uploads/sites/25/2013/12/rat.jpg
5. http://www.ncbi.nlm.nih.gov/gds/
6-7. http://pubchem.ncbi.nlm.nih.gov/search/
8. https://www.youtube.com/watch?v=9mcuIc5O-DE
References:
1. Kawasumi, M ; Nghiem, P. "Chemical Genetics: Elucidating Biological Systems with Small-Molecule Compounds." Journal Of Investigative Dermatology, 2007 Jul, Vol.127(7), pp.1577-1584. http://www.ncbi.nlm.nih.gov/pubmed/17568801
2. "Bone Morphongenetic Protein Inhibitors to Treat Fibrodysplasia Ossificans Progressiva." National Center for Advanced Translational Science. National Institute for Health, n.d. Web. 27 Mar. 2015. <http://www.ncats.nih.gov/research/rare-diseases/trnd/projects/fop.html>.
3. Kaplan, Fs ; Glaser, DL ; Pignolo, Rj ; Shore, Em. "A new era for fibrodysplasia ossificans progressiva: a druggable target for the second skeleton." Expert Opinion On Biological Therapy, 2007 May, Vol.7(5), pp.705-712. http://www.ncbi.nlm.nih.gov/pubmed/17477807
4. "Fibrodysplasia Ossificans Treatment & Management." Fibrodysplasia Ossificans Treatment & Management. WebMD, n.d. Web. 27 Mar. 2015. <http://emedicine.medscape.com/article/1112501-treatment>.
5. "What Are 'model Organisms'?" What Are Model Organisms? Wellcome Trust, n.d. Web. 27 Mar. 2015. <http://genome.wellcome.ac.uk/doc_wtd020803.html>.
6. "Daf-1 (gene) - WormBase : Nematode Information Resource." Daf-1 (gene) - WormBase : Nematode Information Resource. National Institute for Health, n.d. Web. 27 Mar. 2015. <http://www.wormbase.org/species/c_elegans/gene/WBGene00000897#0-9e67-10>.
7. "Acvr1 Targeted Allele Detail MGI Mouse (MGI:5471642)." Acvr1 Targeted Allele Detail MGI Mouse (MGI:5471642). Mouse Genome Informatics, n.d. Web. 27 Mar. 2015. <http://www.informatics.jax.org/allele/MGI:5471642>.
8."RNA Interference Silences Cell-Signaling That Leads to Excessive Bone Formation in FOP." RNA Interference Silences Cell-Signaling That Leads to Excessive Bone Formation in FOP. National Institute for Arthritis and Musculoskeletal and Skin Diseases, n.d. Web. 27 Mar. 2015. <http://www.niams.nih.gov/news_and_events/Spotlight_on_Research/2012/rna_and_fop.asp>.
9. "How RNAi Works." University of Massachusetts Medical School. UMass Medical School, n.d. Web. 27 Mar. 2015. <http://www.umassmed.edu/rti/biology/how-rnai-works/>.
10. "GEO Background." NCBI. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015. <http://www.ncbi.nlm.nih.gov/geo/info/profiles.html>.
1. http://cdn.isciencetimes.com/data/images/full/2014/05/16/6508-mouse.jpg
2. http://content62.eol.org/content/2012/09/26/23/54778_orig.jpg
3. http://www.redorbit.com/media/uploads/2012/02/science-021712-001-617x416.jpg
4. http://arlingtonva.s3.amazonaws.com/wp-content/uploads/sites/25/2013/12/rat.jpg
5. http://www.ncbi.nlm.nih.gov/gds/
6-7. http://pubchem.ncbi.nlm.nih.gov/search/
8. https://www.youtube.com/watch?v=9mcuIc5O-DE
References:
1. Kawasumi, M ; Nghiem, P. "Chemical Genetics: Elucidating Biological Systems with Small-Molecule Compounds." Journal Of Investigative Dermatology, 2007 Jul, Vol.127(7), pp.1577-1584. http://www.ncbi.nlm.nih.gov/pubmed/17568801
2. "Bone Morphongenetic Protein Inhibitors to Treat Fibrodysplasia Ossificans Progressiva." National Center for Advanced Translational Science. National Institute for Health, n.d. Web. 27 Mar. 2015. <http://www.ncats.nih.gov/research/rare-diseases/trnd/projects/fop.html>.
3. Kaplan, Fs ; Glaser, DL ; Pignolo, Rj ; Shore, Em. "A new era for fibrodysplasia ossificans progressiva: a druggable target for the second skeleton." Expert Opinion On Biological Therapy, 2007 May, Vol.7(5), pp.705-712. http://www.ncbi.nlm.nih.gov/pubmed/17477807
4. "Fibrodysplasia Ossificans Treatment & Management." Fibrodysplasia Ossificans Treatment & Management. WebMD, n.d. Web. 27 Mar. 2015. <http://emedicine.medscape.com/article/1112501-treatment>.
5. "What Are 'model Organisms'?" What Are Model Organisms? Wellcome Trust, n.d. Web. 27 Mar. 2015. <http://genome.wellcome.ac.uk/doc_wtd020803.html>.
6. "Daf-1 (gene) - WormBase : Nematode Information Resource." Daf-1 (gene) - WormBase : Nematode Information Resource. National Institute for Health, n.d. Web. 27 Mar. 2015. <http://www.wormbase.org/species/c_elegans/gene/WBGene00000897#0-9e67-10>.
7. "Acvr1 Targeted Allele Detail MGI Mouse (MGI:5471642)." Acvr1 Targeted Allele Detail MGI Mouse (MGI:5471642). Mouse Genome Informatics, n.d. Web. 27 Mar. 2015. <http://www.informatics.jax.org/allele/MGI:5471642>.
8."RNA Interference Silences Cell-Signaling That Leads to Excessive Bone Formation in FOP." RNA Interference Silences Cell-Signaling That Leads to Excessive Bone Formation in FOP. National Institute for Arthritis and Musculoskeletal and Skin Diseases, n.d. Web. 27 Mar. 2015. <http://www.niams.nih.gov/news_and_events/Spotlight_on_Research/2012/rna_and_fop.asp>.
9. "How RNAi Works." University of Massachusetts Medical School. UMass Medical School, n.d. Web. 27 Mar. 2015. <http://www.umassmed.edu/rti/biology/how-rnai-works/>.
10. "GEO Background." NCBI. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015. <http://www.ncbi.nlm.nih.gov/geo/info/profiles.html>.