This web page was produced as an assignment for genetics 564, an undergraduate course at UW-Madison
Gene Homology
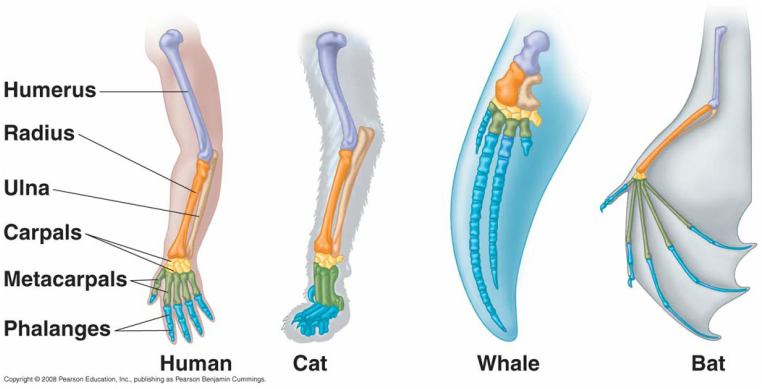
Homology can be defined as similarities between two or more species due to shared ancestry (1,2,3). These similarities can be molecular and/or structural. The image above depicts morphological homology, illustrating how the forelimbs in various mammals with shared ancestry retained similar forelimb skeletal structures throughout evolution. Additionally, homology can be on a molecular level. Sequence homology is the similarity of two DNA sequences or protein sequences between species that have diverged in a speciation event from a common ancestor (3). Gene sequences can change throughout time by accumulating and retaining mutations. In general, species with similar gene sequences tend to be more closely related and have more recently diverged from a speciation event. Genes that are similar between species are said to be conserved. By looking at gene homology in model organisms, this information can help researchers deduce information about evolution, human molecular mechanism or even treatments to diseases.
How Do We Study Homology?
Basic Local Alignment Search Tool, otherwise known as BLAST, is a bioinformatic technique that allows for the comparison between nucleotide or amino acid sequences by aligning them and assessing the similarities (1,3). BLAST is very valuable tool for identifying homologs for a given sequence in many species. BLAST works by taking the input sequence and comparing it to the other sequences in the database to find ones that are the most similar. BLAST was utilized in order to find homologs of ACVR1, which is the disease gene in FOP.
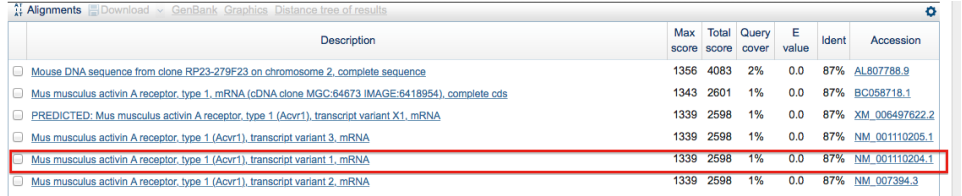
Below is an example of how a mouse ACVR1 gene homolog was found using BLAST. The box in red indicates the homolog found. This demonstrates that the gene in mice is 87 percent identical to the human gene. The last column describes the accession number, which is pertinent because it is the identification number to that specific gene sequence.
Homology can be defined as similarities between two or more species due to shared ancestry (1,2,3). These similarities can be molecular and/or structural. The image above depicts morphological homology, illustrating how the forelimbs in various mammals with shared ancestry retained similar forelimb skeletal structures throughout evolution. Additionally, homology can be on a molecular level. Sequence homology is the similarity of two DNA sequences or protein sequences between species that have diverged in a speciation event from a common ancestor (3). Gene sequences can change throughout time by accumulating and retaining mutations. In general, species with similar gene sequences tend to be more closely related and have more recently diverged from a speciation event. Genes that are similar between species are said to be conserved. By looking at gene homology in model organisms, this information can help researchers deduce information about evolution, human molecular mechanism or even treatments to diseases.
How Do We Study Homology?
Basic Local Alignment Search Tool, otherwise known as BLAST, is a bioinformatic technique that allows for the comparison between nucleotide or amino acid sequences by aligning them and assessing the similarities (1,3). BLAST is very valuable tool for identifying homologs for a given sequence in many species. BLAST works by taking the input sequence and comparing it to the other sequences in the database to find ones that are the most similar. BLAST was utilized in order to find homologs of ACVR1, which is the disease gene in FOP.
Below is an example of how a mouse ACVR1 gene homolog was found using BLAST. The box in red indicates the homolog found. This demonstrates that the gene in mice is 87 percent identical to the human gene. The last column describes the accession number, which is pertinent because it is the identification number to that specific gene sequence.
Gene Homologs to ACVR1 in humans
Humans (Homo sapiens) Dog (Canis lupus familiaris) Mouse (Mus musculus)
activin A receptor, type I (ACVR1) activin A receptor, type 1 (ACVR1) activin A receptor, type I (ACVR1)
Accession #: NM_001105.4 Accession #: XM_549615.4 Accession #: NM_001110204.1 Location: Chromosome 2 95% Identical 87% Identical
2q23-q24
Chimpanzee (Pan troglodytes) Cat (Felis catus) Chicken (Gallus gallus)
activin A receptor, type I (ACVR1) activin A receptor, type I (ACVR1) activin A receptor, type I (ACVR1)
Accession #: XM_009443520.1 Accession #: XM_003990788.2 Accession #: NM_2045601.1
99% Identical 94% Identical 84% Identical
Zebrafish (Danio rerio) Fruit Fly (Drosophila melanogaster) Horse (Equus caballus)
activin A receptor, type I like (acvr1) saxaphone (sax) activin A receptor, type I
Accession #: NC_007113.6 Accession #: NT_033778.4 Accession #: NC_009161.2
Identity unknown Identity unknown 96% Identical
Humans (Homo sapiens) Dog (Canis lupus familiaris) Mouse (Mus musculus)
activin A receptor, type I (ACVR1) activin A receptor, type 1 (ACVR1) activin A receptor, type I (ACVR1)
Accession #: NM_001105.4 Accession #: XM_549615.4 Accession #: NM_001110204.1 Location: Chromosome 2 95% Identical 87% Identical
2q23-q24
Chimpanzee (Pan troglodytes) Cat (Felis catus) Chicken (Gallus gallus)
activin A receptor, type I (ACVR1) activin A receptor, type I (ACVR1) activin A receptor, type I (ACVR1)
Accession #: XM_009443520.1 Accession #: XM_003990788.2 Accession #: NM_2045601.1
99% Identical 94% Identical 84% Identical
Zebrafish (Danio rerio) Fruit Fly (Drosophila melanogaster) Horse (Equus caballus)
activin A receptor, type I like (acvr1) saxaphone (sax) activin A receptor, type I
Accession #: NC_007113.6 Accession #: NT_033778.4 Accession #: NC_009161.2
Identity unknown Identity unknown 96% Identical
Analysis:
The homologs obtained exemplify a few main points. The gene is well conserved in these vertebrate species. This information also indicates that this gene has an essential function as shown by the fact that over millions of years, it has mutated very little among species. This makes sense considering this gene helps facilitate bone formation which is necessary for vertebrates. In regards to invertebrates, although the protein is relatively conserved in invertebrates, the nucleotide sequence is not and could not be derived from a BLAST search.
The homologs obtained exemplify a few main points. The gene is well conserved in these vertebrate species. This information also indicates that this gene has an essential function as shown by the fact that over millions of years, it has mutated very little among species. This makes sense considering this gene helps facilitate bone formation which is necessary for vertebrates. In regards to invertebrates, although the protein is relatively conserved in invertebrates, the nucleotide sequence is not and could not be derived from a BLAST search.
Image References:
1.http://www.bio.miami.edu/dana/pix/homologous_forelimbs.jpg
2. http://blast.ncbi.nlm.nih.gov/Blast.cgi
References:
1. "Discover Homologs." National Center for Biotechnology Information. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015. <http://www.ncbi.nlm.nih.gov/homologene>.
2. "Homologous Genes." Homologous Genes. N.p., 2004. Web. 27 Mar. 2015. <http://evolution.berkeley.edu/evolibrary/article/1_0_0/eyes_10>.
3. Cui, Xf ; Vinar, T ; Brejova, B ; Shasha, D ; Li, M. "Homology search for genes". Bioinformatics, 2007 Jul 1, Vol.23(13), pp.I97-I103. http://bioinformatics.oxfordjournals.org/content/23/13/i97.short
1.http://www.bio.miami.edu/dana/pix/homologous_forelimbs.jpg
2. http://blast.ncbi.nlm.nih.gov/Blast.cgi
References:
1. "Discover Homologs." National Center for Biotechnology Information. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015. <http://www.ncbi.nlm.nih.gov/homologene>.
2. "Homologous Genes." Homologous Genes. N.p., 2004. Web. 27 Mar. 2015. <http://evolution.berkeley.edu/evolibrary/article/1_0_0/eyes_10>.
3. Cui, Xf ; Vinar, T ; Brejova, B ; Shasha, D ; Li, M. "Homology search for genes". Bioinformatics, 2007 Jul 1, Vol.23(13), pp.I97-I103. http://bioinformatics.oxfordjournals.org/content/23/13/i97.short